Mathematically modelling a genetic circuit to detect cancer
Project goal: Use synthetic biology to create a genetic circuit that facilitates the expression of chromoproteins dependent on the presence of lung cancer biomarkers within users' breath. This device would take the form of a breathalyzer and serve as a more accessible, noninvasive avenue for screening for lung cancer.
Tech Stack: Python (ODE simulation), Scipy, Numpy, matplotlib
Only care about software? Skip to the "Computational Methods, Results, and Discussion" section.
Abstract (TL;DR for software readers)
Bronchogenic carcinoma (lung cancer) remains a leading cause of cancer‐related mortality, with current diagnostic methods limited by cost and accessibility. In response, this novel, cost‐effective biosensor that uses a synthetic biological circuit engineered in yeast was developed to detect lung cancer‐associated volatile organic compounds (VOCs) early. The circuit employs an AND gate logic whereby 2‐butanone detection via an olfactory receptor induces VP16 expression, and 1‐propanol detection through a GLD1⁺ promoter yields GAL4‐DBD production. The concomitant presence of both biomarkers facilitates the formation of a GAL4‐VP16 complex, thereby activating green chromoprotein expression. A supplementary glucose‐responsive branch, producing blue chromoprotein, serves as a false‐negative indicator. Validation using computational simulations and data analysis confirmed circuit performance with minimal leaky expression. This biosensor represents a promising advance for accessible lung cancer screening, with the potential to improve clinical outcomes through timely diagnosis and intervention. These promising results warrant further experimental validation and clinical investigation in future studies.
Introduction
Bronchogenic carcinoma, commonly known as lung cancer, refers to tumors originating within the bronchi or lung parenchyma [1]. In Canada, approximately ¼ of cancer-related deaths are attributed to lung cancer, and 98% of lung cancer diagnoses occur in people over 50 [2]. Lung cancer is difficult to diagnose at early stages as patients often remain asymptomatic until the cancer progresses, making early detection difficult [3].
Current lung cancer diagnosis involves imaging tests, laboratory analyses, and specialized procedures; however, they are not readily available to individuals of different socio-economic/geographic backgrounds due to high costs [4]. In response, many nanomaterial-based sensors have been reported for lung cancer biomarkers, but none are commercially available [5].
Lung cancer gives off volatile organic compounds (VOCs) as a result of oxidative stress in the lungs [6]. The VOCs given off by lung cancer cells include alkanes and identifiable biomarkers [6]. While no study considered a single VOC as sufficient for diagnosis, researchers agree that a set of substances and their chemometric fingerprint is required for an unambiguous diagnosis.[7]. Five different studies identified by Saalberg et al. [8], have hypothesized 2-Butanone and 1-Propanol as very strong biomarkers for lung cancer [7]
To address the problem of diagnostic inaccessibility, the proposed solution utilizes two substances and their chemometric fingerprint for a more robust diagnosis and early detection of lung cancer via a color-changing breathalyzer. This solution leverages a biological AND gate that activates green chromoprotein through the use of olfactory receptors and biological components in yeast, which are able to detect 2-Butanone and 1-Propanol.
The proposed solution consists of three branches for the detection of three distinct chemicals, and expresses two different colors through chromoprotein expression. In one branch, 2-Butanone is detected using an olfactory receptor, Olfr1258 [8]. Olfactory receptors, often described as chemical noses, give off small molecules when they detect a specific chemical [9]. Olfr1258 releases GPCR, which in turn positively induces the promoter. Once the promoter is induced, the branch is activated, resulting in the expression of VP16 [10]. Another branch is used to detect 1-Propanol. GLD1+ promoter is directly induced by 1-Propanol, resulting in the activation of the branch and expression of the GAL4-DBD [11]. However, GLD1+ is simultaneously repressed by glucose, and while glucose is not commonly found in a fasted breath [11], a third branch was to indicate a false negative. The HXT-1 Promoter is used as it is induced by glucose [12], resulting in the expression of blue chromoprotein. Blue chromoprotein is thus used to signify a false negative. In the case that both 1-Propanol and 2-Butanone are present in the breath, it has been established that both the Gal4-DBD and VP16 will be produced by the circuit. If the 2 molecules are present, the DNA-binding domain of Gal4 fuses with the activation domain of VP16, forming Gal4-VP16 [13]. The complex induces a promoter which activates the RBS and coding sequence, expresses the GLP chromoprotein, which is fluorescent green in colour, indicating the presence of both 2-Butanone and 1-Propanol, both significant biomarkers of lung cancer.
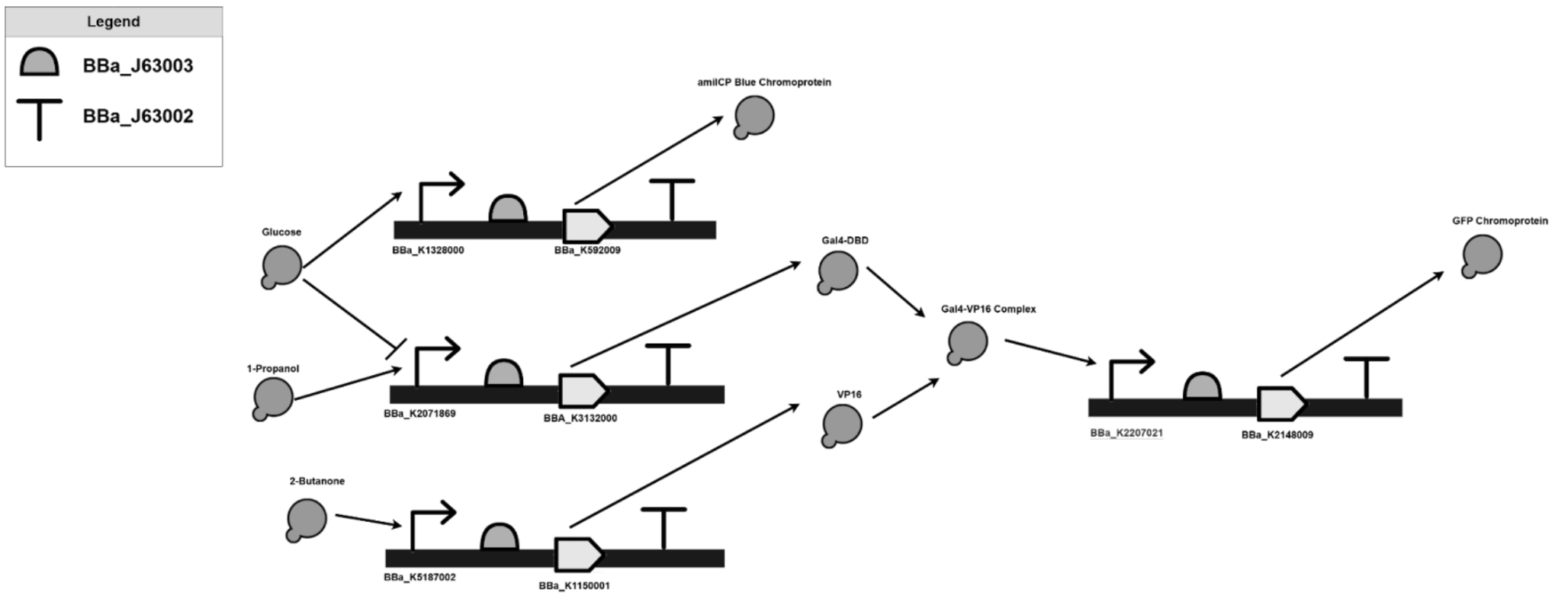
Such a system has the potential to facilitate early detection, improving clinical outcomes through timely intervention, and allowing for more accessible diagnosis solutions.
Materials and In-Vitro Interpretations
The Proposed Circuit Construction
The parts for the proposed biological circuits are listed in Table 1, are all “RFC [10]” “RFC [12]”, or “RFC [1000]” compatible and are eligible for BioBrick Assembly [14]. The biological parts come in a plasmid form and will need to be digested using restriction enzymes to prepare for insertion. A reaction using EcoRl and SpeI restriction enzymes should be individually completed for each.
To amplify each part, PCR is performed using the reagents (see Table 2), abiding by the protocol described in [15]. Gel electrophoresis is used to analyze the PCR product, verifying the success and presence of the target DNA sequence in the sample [16]. The reagents listed in Table 2 are used in the procedure, following the protocols described in [17]. The remaining PCR product is purified using the PureLink PCR Purification Kit.
Following BioBrick assembly procedure, EcoRI and SpeI are first used to cleave the desired part at its BioBrick upstream region. EcoRI cuts at 5'-GAATTC-3' and SpeI cuts at 5'-A/CTAGT-3', leaving sticky ends. Next, XbaI and PstI cleave the desired downstream sites at 5'-TCTAGA-3' and 5'-CTGCAG-3', respectively [18]. Then, EcoRI and PstI are used to cleave the destination plasmid (pSKB3K3). Finally, the corresponding sticky ends join up so the final composite plasmid consists of cuts in the order of EcoRI, XbaI, upstream part region, downstream part region, SpeI, and PstI [19]. These steps will be repeated for each BioBrick part, as these parts will be added one at a time.
Using BioBrick assembly, each part that has undergone PCR amplification is inserted one at a time in the destination plasmid utilizing T4 DNA ligase. The parts of the biological circuit are inserted (from 5 ’ to 3’). Furthermore, a control tube is constructed using the same procedure, and replacing ligase with nuclease-free water.
Expected Results and Interpretations:
Group 1, a negative control, is established by including a group of yeast that is not modified by the proposed biological circuit. The negative control should express no chromoproteins in the presence of either 2-Butanone, 1-Propanol and/or Glucose. Group 2, a positive control, is designed by having a yeast cell that continuously expresses a positive signal regardless of the presence of 2-Butanone, 1-Propanol and/or Glucose. This is done by having GFP readily available in the circuit at all times.
Groups A-E are designed by integrating the proposed biological circuit into the yeast cell. Group A is absent from both biomarkers and glucose; no chromoprotein expression is predicted for this group, ensuring baseline repression. Group B contains 1-Propanol but not 2-Butanone and glucose to test the efficacy of the propanal detecting branch. The concentration value is selected to be within the accepted pathogenic range. It is predicted that this would result in the expression of the Gal4-DBD molecule. Group C tests the 2-Butanone detention branch by containing 2-Butanone but not 1-Propanol and glucose. It is expected that Group C would allow for the formation of the VP16 molecule, and no chromoprotein expression. Group D would test the AND logic of the circuit by containing 2-Butanone and 1-Propanol. Group D would then express the green chromoprotein, as well as the Gal4-VP16. Finally, Group E is there to test the effectiveness of the glucose-detecting branch; it will contain the same values as Group D in addition to glucose. It is predicted that the green chromoprotein will be turned off, and instead, the blue chromoprotein will be expressed.
Blue chromoprotein expression in any group except Group C indicates a flaw with the detection of glucose. If green chromoprotein expression is detected in Group 1 or Groups A-D, it suggests leaky expression or a flaw in the AND gate. Similarly, if Group E is absent from the expression of the green chromoprotein, an issue with the AND gate is predicted.
Computational Methods, Results, and Discussion
This system was initially modelled in MATLAB SimBiology, but was subsequently translated into a custom, object-oriented Python architecture for freedom of analysis. This computational environment enabled a departure from GUI-based assumptions to build a highly transparent, physics-informed numerical model.
1. The Mathematical Framework (The Physics Engine)
To move away from GUI-dependent software like SimBiology, the entire biological circuit had to be mathematically defined using a system of Ordinary Differential Equations (ODEs). This required explicitly tracking the concentration of mRNAs, proteins, and complexes over time based on physical and chemical kinetics:
-
Transcription (Hill Equations): To model promoter activation, cooperative Hill equations were utilized. For example, the transcription of the GAL4-DBD mRNA driven by 1-propanol activation (and repressed by glucose) was modeled as:
(Assuming a global activation threshold
K=50, a Hill coefficientH=2, and a transcription rate of50 bp/snormalized for the cell nucleus volume). -
Translation (Mass Action): Protein translation was modeled via mass action kinetics directly dependent on mRNA molarity:
-
Complexation: The formation of the transcription factor complex acting as the AND gate (GAL4-DBD fusing with VP16) was modeled using an association rate (
10^9 M^-1 s^-1) and dissociation rate (10 s^-1): -
Decay Kinetics: Every biological species in the simulation was subject to a continuous loss rate based on its half-life:
2. Software Architecture: Object-Oriented ODE Solving
Rather than writing a flat, procedural script, the Python architecture was designed using object-oriented principles to keep the biological branches highly modular.
A GeneCircuitODE class managed the system state, while dataclasses were utilized to strictly type and store the biological constants (mRNA half-lives, degradation rates, and cell volume). To optimize execution speed, the mathematical framework was vectorized using numpy arrays.
The core physics engine relied on scipy.integrate.solve_ivp utilizing the explicit Runge-Kutta method (RK45) to integrate the system over a 3600-second timeframe, with a built-in fallback to the Radau solver for stiff equations:
class GeneCircuitODE:
"""Encapsulates the 9-species ODE system for the yeast biosensor."""
def __init__(
self,
inputs: InputConcentrations,
constants: BiologicalConstants | None = None,
):
self.inputs = inputs
self.c = constants or BiologicalConstants()
self.idx = SpeciesIndex()
def rhs(self, t: float, y: np.ndarray) -> np.ndarray:
"""Right-hand side of the ODE system."""
# Clamp to non-negative to handle numerical noise
y = np.maximum(y, 0.0)
c = self.c
inp = self.inputs
dy = np.zeros(self.idx.NUM_SPECIES)
# Unpack state
mRNA_VP16 = y[0]
VP16 = y[1]
mRNA_Gal4DBD = y[2]
Gal4DBD = y[3]
Gal4VP16 = y[4]
mRNA_GFP = y[5]
GFP = y[6]
mRNA_amilCP = y[7]
amilCP = y[8]
# Complexation terms (shared)
formation = complex_formation_rate(Gal4DBD, VP16, c.k_on)
dissociation = complex_dissociation_rate(Gal4VP16, c.k_off)
# Branch 1: Butanone → VP16
dy[0] = hill_activation(inp.butanone, c.K, c.H, c.k_trsc) - decay_rate(mRNA_VP16, c.mRNA_decay)
dy[1] = translation_rate(mRNA_VP16, c.k_trans) - decay_rate(VP16, c.protein_decay) - formation + dissociation
# Branch 2: Propanol (repressed by glucose) → Gal4DBD
dy[2] = (
hill_activation(inp.propanol, c.K, c.H, c.k_trsc)
* hill_repression(inp.glucose, c.K_glucose, c.H)
- decay_rate(mRNA_Gal4DBD, c.mRNA_decay)
)
dy[3] = translation_rate(mRNA_Gal4DBD, c.k_trans) - decay_rate(Gal4DBD, c.protein_decay) - formation + dissociation
# Branch 3: Gal4VP16 complex → GFP
dy[4] = formation - dissociation - decay_rate(Gal4VP16, c.protein_decay)
dy[5] = hill_activation(Gal4VP16, c.K, c.H, c.k_trsc) - decay_rate(mRNA_GFP, c.mRNA_decay)
dy[6] = translation_rate(mRNA_GFP, c.k_trans) - decay_rate(GFP, c.protein_decay)
# Branch 4: Glucose → amilCP
dy[7] = hill_activation(inp.glucose, c.K_glucose, c.H, c.k_trsc) - decay_rate(mRNA_amilCP, c.mRNA_decay)
dy[8] = translation_rate(mRNA_amilCP, c.k_trans) - decay_rate(amilCP, c.protein_decay)
return dy
def simulate(
self,
t_span: tuple[float, float] = (0, 3600),
y0: np.ndarray | None = None,
method: str = "RK45",
n_points: int = 1000,
):
"""Run the simulation and return the solve_ivp result."""
if y0 is None:
y0 = np.zeros(SpeciesIndex.NUM_SPECIES)
t_eval = np.linspace(t_span[0], t_span[1], n_points)
sol = solve_ivp(
self.rhs,
t_span,
y0,
method=method,
t_eval=t_eval,
rtol=1e-8,
atol=1e-10,
dense_output=True,
)
if not sol.success:
# Fall back to stiff solver
sol = solve_ivp(
self.rhs,
t_span,
y0,
method="Radau",
t_eval=t_eval,
rtol=1e-8,
atol=1e-10,
)
return solThis architecture provides total transparency into the mathematical state of the biosensor at any given millisecond, something that "black-box" simulation software often obscures.
Simulation Results:
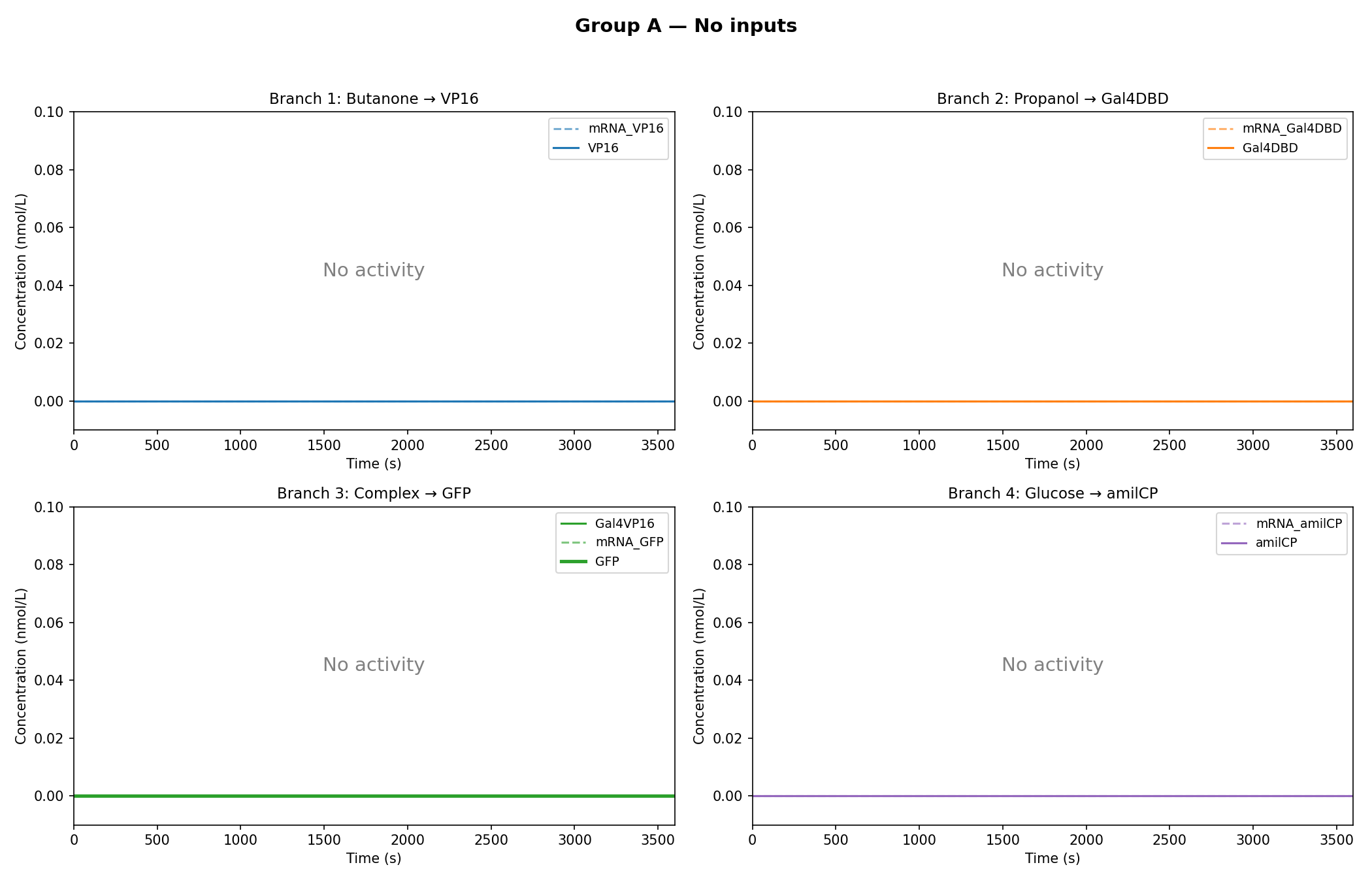
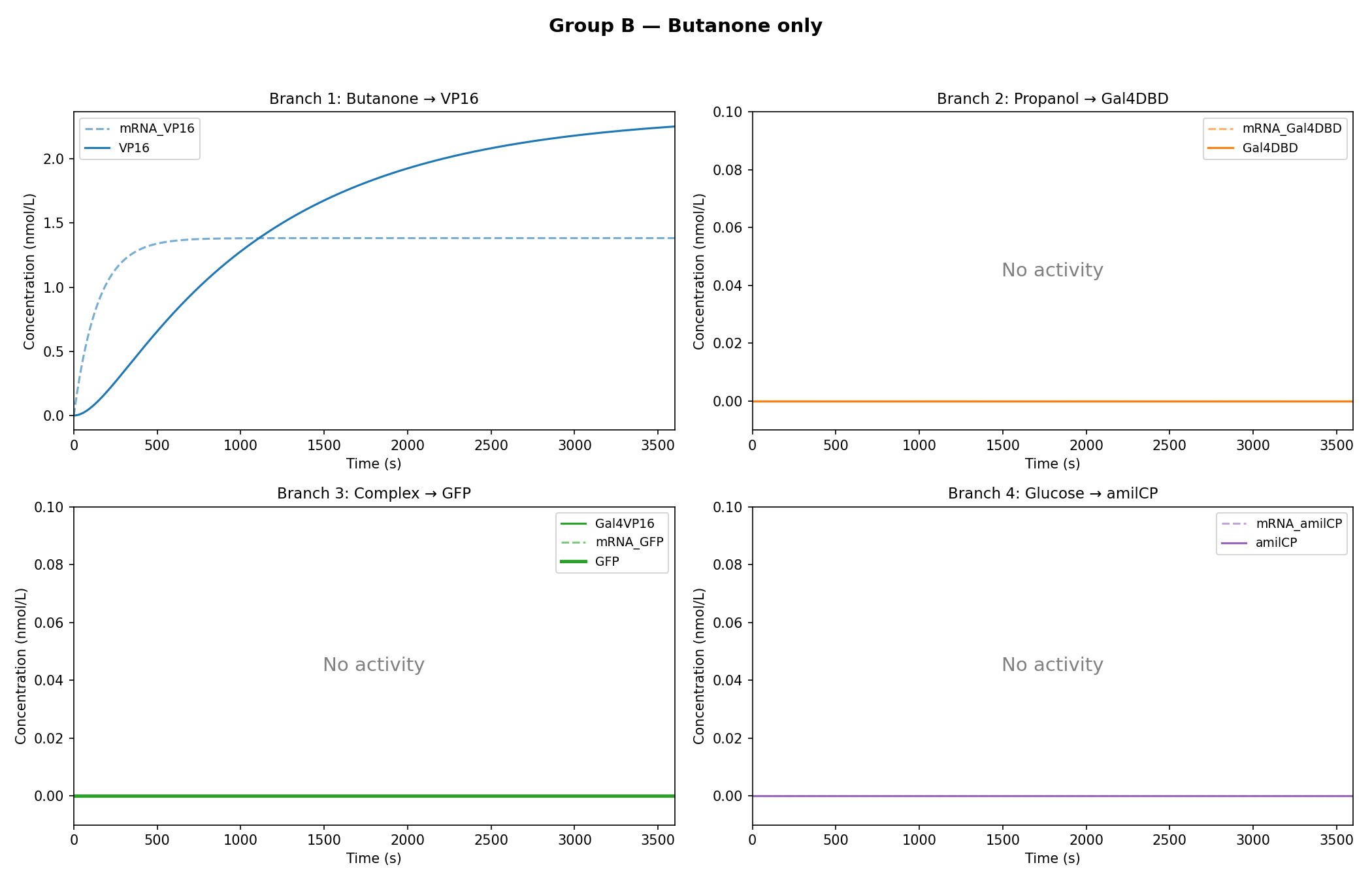
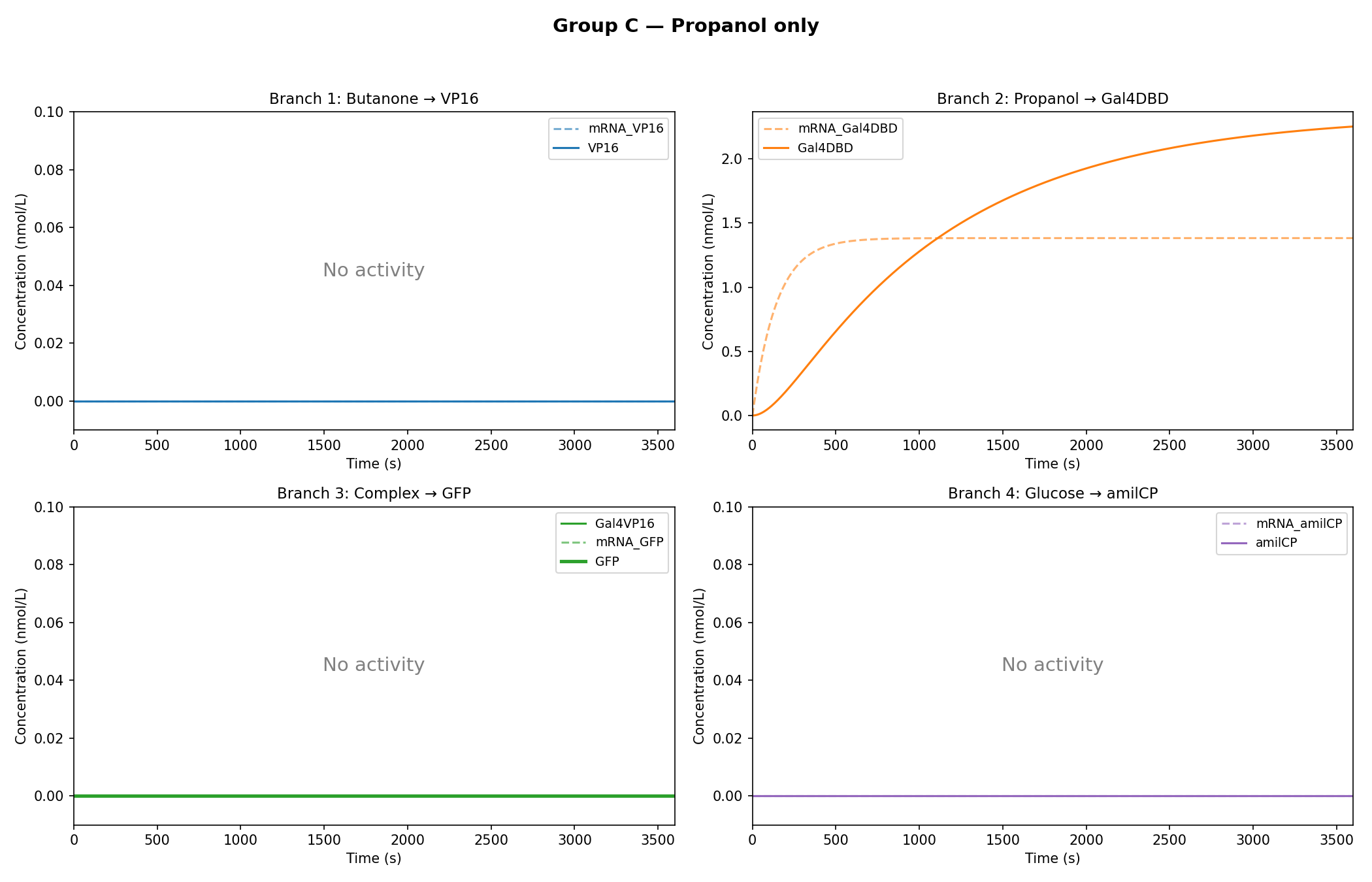
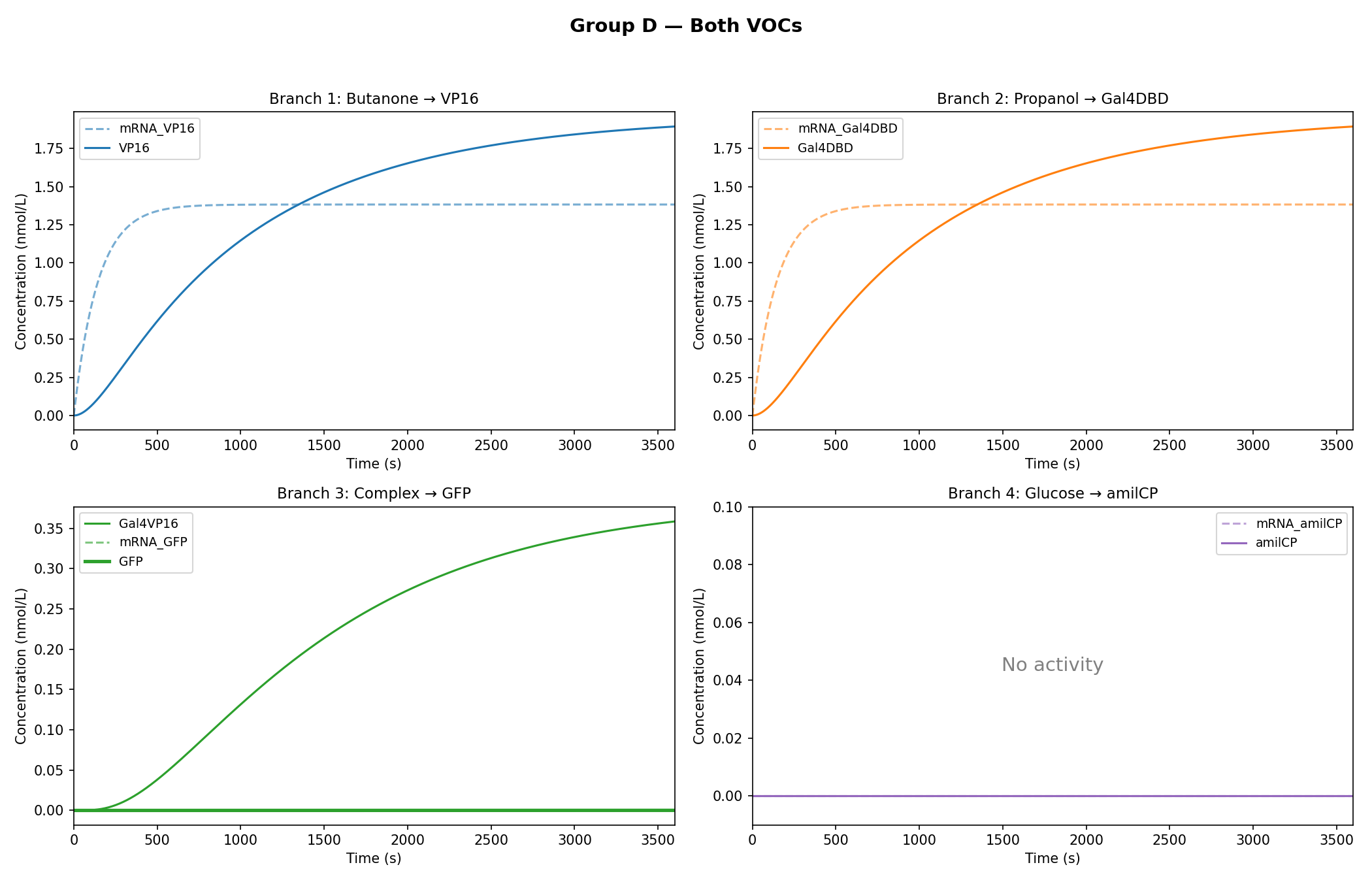
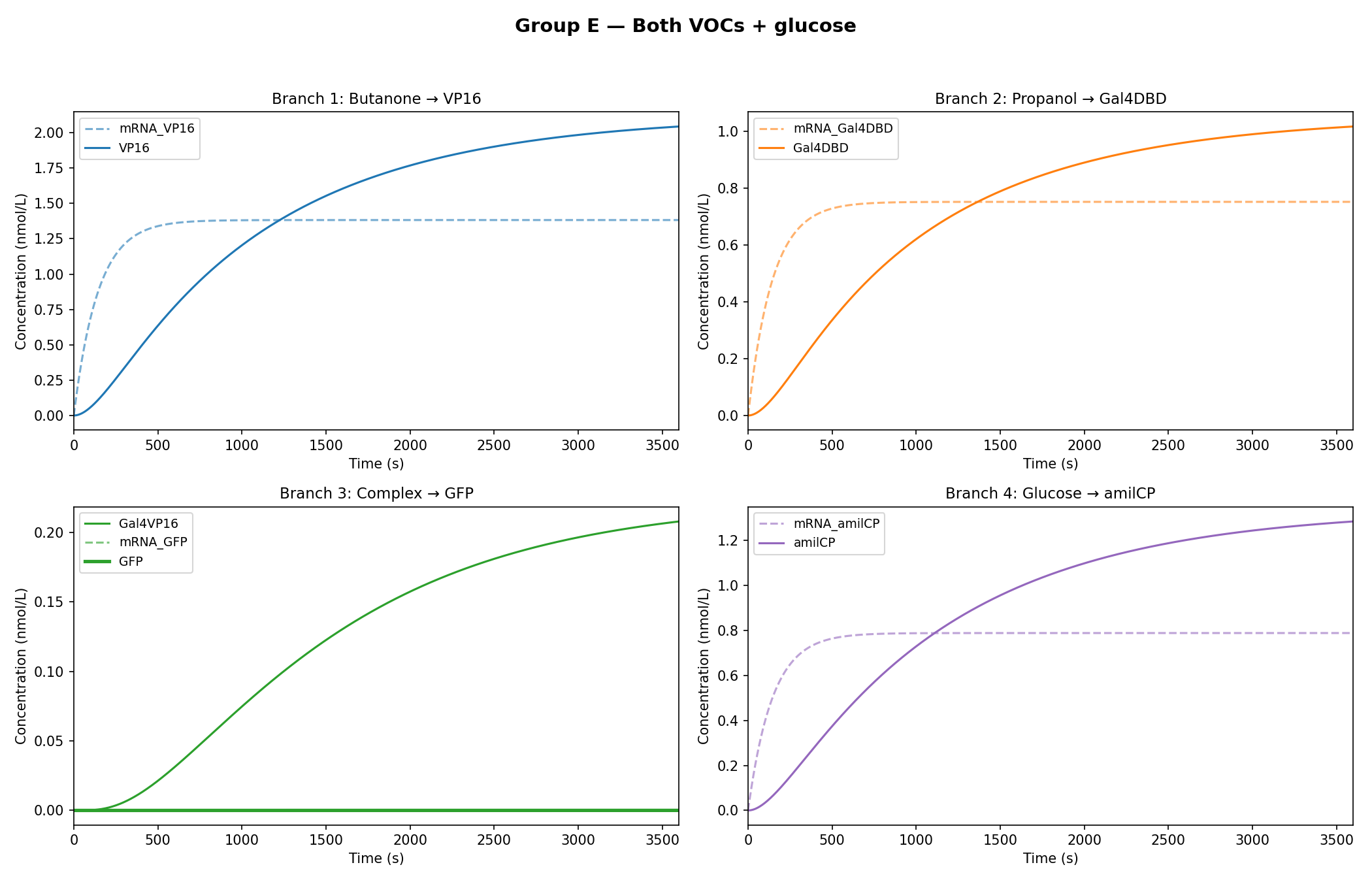
As observed in the figures above, all experimental groups functioned as expected based on the established AND-gate logic.
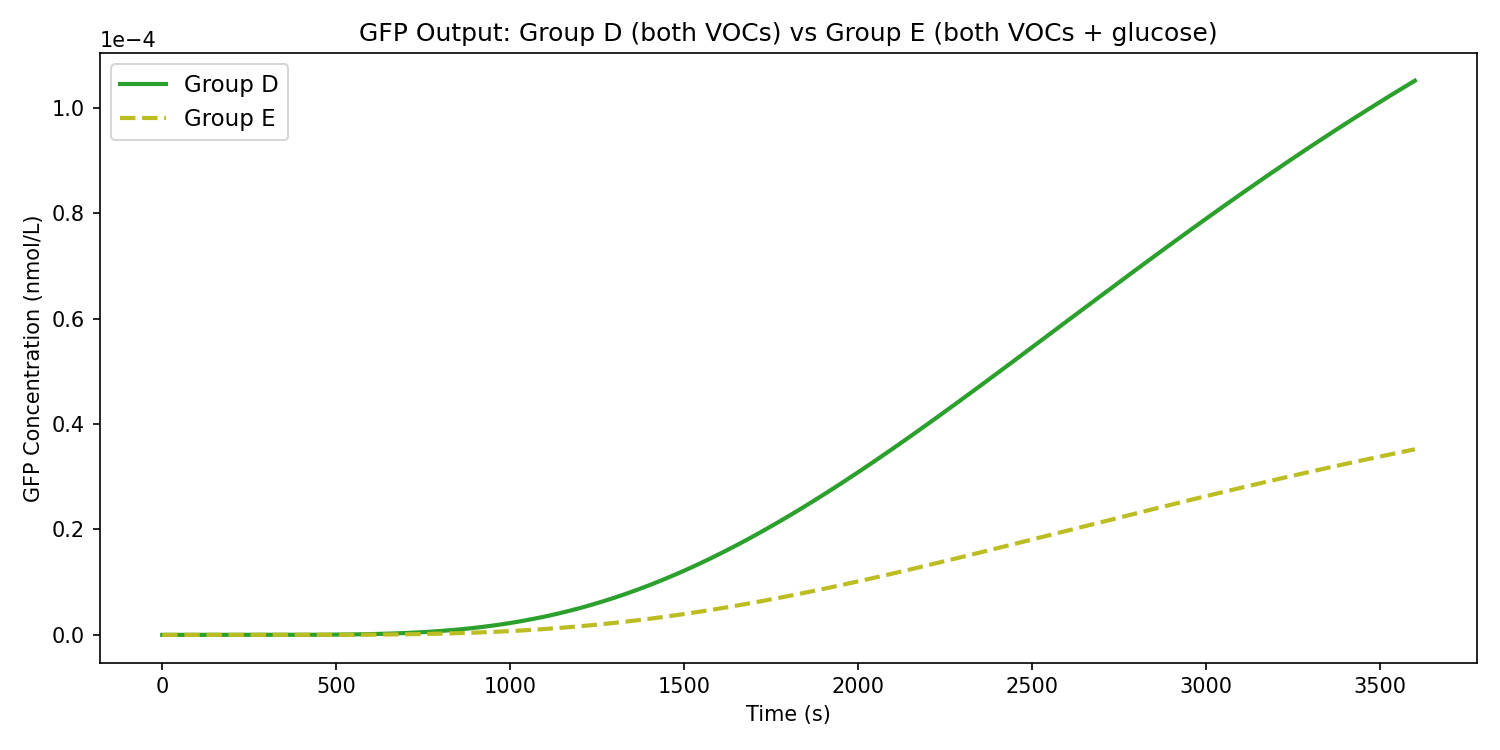
Additionally, it was observed that glucose dampens the expression of GFP, rather than completely inhibiting it.
While the qualitative logic of the circuit held true (Group D correctly formed the GAL4-VP16 complex, and Group E successfully activated the amilCP false-negative indicator), the rigorous mathematical constraints of the Python model exposed critical physical limitations in the biological design.
Engineering Reflections & Biological Limitations:
By keeping the kinetic parameters strictly tied to realistic biological approximations (such as a global activation threshold of K=50), the simulation revealed two major design flaws that would need addressing before in-vitro synthesis:
The Threshold Mismatch (GFP Starvation): The model demonstrated that the GAL4-VP16 complex only reaches a steady state of roughly 0.35 nmol/L. Because this concentration is drastically lower than the K=50 threshold required to fully induce the UAS-GAL promoter, the math dictates that the promoter remains effectively starved, resulting in a microscopic expression of GFP. In a wet lab setting, the circuit requires a significantly stronger promoter for the VOC branches, a cascading biological amplification, or a novel UAS-GAL site with a lower dissociation constant.
Weak Repressor: In Group E, the circuit's ability to respond to the presence of glucose to block the production of the green chromoprotein was tested. Glucose is intended as a “kill-switch” for the 1-propanol branch. However, with the threshold of K=50, this naturally occurring concentration of glucose proved to be mathematically insufficient. The current repressor dampens the branch, allowing Gal4-DBD to continue leaking into the system. In a real clinical diagnostic tool, this increases the risk of false positives, introducing a level of confusion between the appropriate ratio of GFP and amilCP in the activated circuit. Identifying these threshold mismatches computationally saves immense amounts of time and resources, allowing for the precise re-engineering of promoter sensitivities before a single plasmid is ever pipetted in the lab.
Conclusion
The proposed solution presents a non-invasive approach to detecting early-stage lung cancer through targeting two VOCs found in the breath of lung cancer patients. 1-propanol and 2-butanone were selected due to high confidence levels. By translating the biological architecture into a custom Python simulation, we modeled the system's ability to express GFP in the presence of both VOCs under ideal fasted conditions. Furthermore, to prevent false negatives in the presence of glucose, amilCP is expressed and overrides GFP expression.
When analyzing the simulation, it was determined that GFP concentration undergoes significant increase when both biomarkers are present in fasted conditions, thus confirming that the dual-input 1-propanol - 2-butanone system can successfully express GFP through the formation of the GAL4-VP16 complex as expected. In this case, no glucose is present in the breath, and therefore, the amilCP concentration is zero. The model demonstrated that in the scenario that glucose is present in the breath, the 1-propanol branch is repressed (but not completely inhibited), reducing the formation of the GAL4-VP16 complex, which in turn reduces the expression of GFP. Instead, the glucose branch of the circuit is prioritized, causing amilCP to be expressed which rules test results as inconclusive for presence of lung cancer.
Appendix
Table 1: Biological parts used in the circuit.
| Part Name | Type | Function | Supplier |
|---|---|---|---|
| Branch One: 2-Butanone Detection | |||
| Saccharomyces cerevisiae | Fungi | Acts as a host for DNA expression; circuit is introduced via plasmid. | Fisher Scientific [20] |
| BBa_J63002 | Terminator | Signals the end of transcription. | iGEM [21] |
| BBa_K5187002 | Promoter | Positively induced by 2-butanone. | iGEM [22] |
| BBa_J63003 | RBS | Ensures that ribosomes bind to mRNA for translation. | iGEM [21] |
| BBa_K1150001 | CDS for VP16 | Coding sequence for VP16, an intermediate protein required for GFP expression. | iGEM [10] |
| Branch Two: 1-Propanol Detection | |||
| Saccharomyces cerevisiae | Fungi | Acts as a host for DNA expression. | Fisher Scientific [20] |
| BBa_K2071869 | Promoter | Induced by 1-propanol but repressed by glucose. | PubMed [11] |
| BBa_J63003 | RBS | Ensures that ribosomes bind to mRNA. | iGEM [21] |
| BBa_J63002 | Terminator | Signals the end of transcription. | iGEM [21] |
| BBa_K3132000 | CDS for Gal4-DBD | Encodes Gal4-DBD, part of the complex needed for GFP expression. | iGEM [23] |
| Branch Three: Glucose Detection | |||
| Saccharomyces cerevisiae | Fungi | Acts as a host for DNA expression. | Fisher Scientific [20] |
| BBa_J63003 | RBS | Ensures that ribosomes bind to mRNA. | iGEM [21] |
| BBa_J63002 | Terminator | Signals the end of transcription. | iGEM [21] |
| BBa_K1328000 | Promoter | Induced by high glucose concentrations, leading to blue chromoprotein expression. | iGEM [12] |
| BBa_K592009 | CDS for amilCP | Encodes for the expression of amilCP (blue chromoprotein). | iGEM [24] |
| Branch Four: Chromoprotein Expression | |||
| BBa_K2148009 | CDS | Codes for the production of the GFP chromoprotein. | iGEM [25] |
| BBa_K2207021 | Promoter | Induced by Gal4-VP16 complex. | iGEM [26] |
| BBa_J63002 | Terminator | Signals the end of transcription. | iGEM [21] |
| BBa_J63003 | RBS | Ensures that ribosomes bind to mRNA. | iGEM [21] |
Table 2: List of reagents, their intended use, and their supplier.
| Reagent | Use | Supplier |
|---|---|---|
| Culturing & Plasmids | ||
| Kanamycin | Antibiotic for culturing cells. | Millipore Sigma [27] |
| kanMX module | Provides resistance to antibiotic. | Addgene [28] |
| pSB3K3 | Plasmid where the biological circuit will be inserted. | Addgene [29] |
| Digestion of Biological Parts | ||
| EcoRI | Restriction enzyme to digest biological parts, used for insertion. | New England Biolabs [30] |
| SpeI | Restriction enzyme to digest biological parts, used for insertion. | Fisher Scientific [31] |
| XbaI | Restriction enzyme to digest biological parts, used for insertion. | Fisher Scientific [32] |
| PstI | Restriction enzyme to digest biological parts, used for insertion. | Fisher Scientific [33] |
| Amplification and Purification | ||
| Taq Polymerase | Amplifies the target DNA sequence in PCR. | Qiagen [34] |
| Forward and Reverse Primers | Flank the target sequence of the DNA for amplification/cloning. | ThermoFisher [35] |
| MgCl₂ | Enhances the activity of Taq Polymerase, aiding in amplification. | ThermoFisher [36] |
| DNA Template | Contains the target DNA sequence to be amplified. | Qiagen [34] |
| Four dNTPs | Act as the building blocks for replication, creating new DNA strands. | ThermoFisher [37] |
| Buffer | Maintains the pH of the reaction mixture. | ThermoFisher [38] |
| Agarose Gel | Medium for gel electrophoresis to separate DNA segments. | Fisher Scientific [39] |
| Gel stain | Helps to visualize the DNA during gel electrophoresis. | ThermoFisher [40] |
| Loading dye | Allows visualization of sample migration during gel electrophoresis. | ThermoFisher [41] |
| PureLink PCR Purification Kit | Purifies DNA samples. | ThermoFisher [42] |
| Digestion of Destination Plasmid | ||
| EcoRI | Restriction enzyme to digest biological parts, used for prefix insertion. | New England Biolabs [30] |
| SpeI | Restriction enzyme to digest biological parts, used as part of suffix sequence. | Fisher Scientific [31] |
| Preparation of Insert | ||
| T4 DNA Ligase | Assists in insertion of amplified biological parts into destination plasmid. | ThermoFisher [43] |
| T4 DNA Ligase Buffer | Buffer for ligation reaction. | New England Biolabs [44] |
| Nuclease Free Water | Replaces the ligase in control reaction. | ThermoFisher [45] |
| LB Agar Plates | Act as a growth medium for E. coli colonies. | ThermoFisher [46] |
References
[1] F. Siddiqui and A. H. Siddiqui, “Lung Cancer,” PubMed, 2021. https://pubmed.ncbi.nlm.nih.gov/29493979/ [2] Canadian Cancer Society, “About Lung Cancer: Lung Cancer in Canada | Canadian Lung Association,” www.lung.ca, 2023. https://www.lung.ca/lung-health/lung-diseases/lung-cancer/about-lung-cancer-lung-cancer-canada [3] J. Ning et al., “Early diagnosis of lung cancer: which is the optimal choice?,” Aging (Albany NY), vol. 13, no. 4, pp. 6214–6227, Feb. 2021, doi: https://doi.org/10.18632/aging.202504. [4] Mayo Clinic, “Lung Cancer - Diagnosis and Treatment - Mayo Clinic,” Mayoclinic.org, Mar. 22, 2022. https://www.mayoclinic.org/diseases-conditions/lung-cancer/diagnosis-treatment/drc-20374627 [5] Raja Chinnappan et al., “Emerging Biosensing Methods to Monitor Lung Cancer Biomarkers in Biological Samples: A Comprehensive Review,” Cancers, vol. 15, no. 13, pp. 3414–3414, Jun. 2023, doi: https://doi.org/10.3390/cancers15133414. [6] M. Phillips et al., “Volatile organic compounds in breath as markers of lung cancer: a cross-sectional study,” The Lancet, vol. 353, no. 9168, pp. 1930–1933, Jun. 1999, doi: https://doi.org/10.1016/s0140-6736(98)07552-7. [7] Y. Saalberg and M. Wolff, “VOC breath biomarkers in lung cancer,” Clinica Chimica Acta, vol. 459, pp. 5–9, Aug. 2016, doi: https://doi.org/10.1016/j.cca.2016.05.013. [8] “Part:BBa_K2329004 - parts.igem.org,” Igem.org, 2017. https://parts.igem.org/Part:BBa_K2329004 [9] I. Glezer and B. Malnic, “Olfactory receptor function,” Handbook of Clinical Neurology, vol. 164, pp. 67–78, 2019, doi: https://doi.org/10.1016/B978-0-444-63855-7.00005-8. [10] “Part:BBa_K1150001 - parts.igem.org,” Igem.org, 2019. https://parts.igem.org/Part:BBa_K1150001 [11] T. Matsuzawa, T. Ohashi, A. Hosomi, N. Tanaka, H. Tohda, and K. Takegawa, “The gld1+ gene encoding glycerol dehydrogenase is required for glycerol metabolism in Schizosaccharomyces pombe,” Applied microbiology and biotechnology, vol. 87, no. 2, pp. 715–27, Jun. 2010, doi: https://doi.org/10.1007/s00253-010-2586-3. [12] “Part:BBa_K1328000 - parts.igem.org,” parts.igem.org. https://parts.igem.org/Part:BBa_K1328000 [13] G. A. Jacob and D. S. Luse, “GAL4-VP16 Stimulates Two RNA Polymerase II Promoters Primarily at the Preinitiation Complex Assembly Step,” Gene Expression, vol. 5, no. 3, p. 193, Sep. 2018. Available: https://pmc.ncbi.nlm.nih.gov/articles/PMC6138014/ [14] “iGEM Technology,” technology.igem.org. https://technology.igem.org/assembly/biobrick [15] Merck, "NMR Chemical Shifts of Impurities," Merck, vol. 1, no. 1, 2025. Available: https://www.sigmaaldrich.com/MX/en/technical-documents/technical-article/genomics/cloning-and-expression/blue-white-screening [16] A. Carrillo, "Gel Electrophoresis Steps," Azure Biosystems, Jan. 18, 2023. Available: https://azurebiosystems.com/blog/gel-electrophoresis-steps/ [17] "PureLink ® PCR Purification Kit For rapid, efficient purification of PCR products." Available: https://assets.thermofisher.com/TFS-Assets/LSG/manuals/purelink_pcr_man.pdf [18] "DNA/Assembly - parts.igem.org," Igem.org, 2025. Available: https://parts.igem.org/DNA/Assembly [19] New England Biolabs, "BioBrick Assembly Kit," Neb.com, 2021. Available: https://www.neb.com/en-ca/products/e0546-biobrick-assembly-kit [20] Microbiologics, "Saccharomyces cerevisiae ATCC™ 9763™," Fisher Scientific, 2025. Available: https://www.fishersci.com/shop/products/microbiologics-i-saccharomyces-cerevisiae-i-atcc-9763-3/p-4395662 [21] iGEM Parts Registry, "Part:BBa_J63003," 2006. Available: https://parts.igem.org/Part:BBa_J63003 [22] iGEM Parts Registry, "Part:BBa_K5187002," 2024. Available: https://parts.igem.org/Part:BBa_K5187002 [23] iGEM Parts Registry, "Part:BBa_K3132000," 2019. Available: https://parts.igem.org/Part:BBa_K3132000 [24] iGEM Parts Registry, "Part:BBa_K592009," 2011. Available: https://parts.igem.org/Part:BBa_K592009 [25] iGEM Parts Registry, "Part:BBa_K2148009," 2016. Available: https://parts.igem.org/Part:BBa_K2148009 [26] iGEM Parts Registry, "Part:BBa_K2207021," 2017. Available: https://parts.igem.org/Part:BBa_K2207021 [27] Sigma-Aldrich, "Kanamycin sulfate from Streptomyces kanamyceticus," Sigma-Aldrich, 2025. Available: https://www.sigmaaldrich.com/CA/en/product/sigma/k1377 [28] Addgene, "pFA6a-6xGLY-3xFLAG-kanMX6," Addgene, 2025. Available: https://www.addgene.org/20754/ [29] Addgene, "pSB3K3 containing mRFP1," Addgene, 2025. Available: https://www.addgene.org/118083/ [30] New England Biolabs, "EcoRI," New England Biolabs, 2025. Available: https://www.neb.com/en/products/r0101-ecori [31] Thermo Fisher Scientific, "BcuI (SpeI) (10 U/μL)," Thermo Fisher Scientific, 2025. Available: https://www.fishersci.ca/shop/products/bcui-spei/ferer1251 [32] Thermo Fisher Scientific, "XbaI, HC (50 U/μL)," Thermo Fisher Scientific, 2025. Available: https://www.fishersci.ca/shop/products/xbai-2/FERER0683#?keyword=XbaI [33] Invitrogen, "Anza™ 23 PstI," Thermo Fisher Scientific, 2025. Available: https://www.fishersci.ca/shop/products/anza-23-psti-2/IVGN0236 [34] QIAGEN, "Taq DNA Polymerase," QIAGEN, 2025. Available: https://www.qiagen.com/us/products/discovery-and-translational-research/enzymes-for-molecular-biology/taq-dna-polymerase [35] Thermo Fisher Scientific, "High-Quality Modified Nucleotides for Research," Thermo Fisher Scientific, 2025. Available: https://www.thermofisher.com/ie/en/home/life-science/oligonucleotides-primers-probes-genes/nucleotides.html [36] Thermo Fisher Scientific, "MgCl₂ (magnesium chloride) (25 mM)," Thermo Fisher Scientific, 2025. Available: https://www.thermofisher.com/order/catalog/product/R0971 [37] Thermo Fisher Scientific, "dNTP Mix (10 mM each)," Thermo Fisher Scientific, 2025. Available: https://www.thermofisher.com/order/catalog/product/R0193 [38] Invitrogen, "Taq DNA Polymerase PCR Buffer (10X)," Thermo Fisher Scientific, 2025. Available: https://www.thermofisher.com/order/catalog/product/18067017 [39] Fisher Scientific, "Agarose Electrophoresis Gels," Fisher Scientific, 2025. Available: https://www.fishersci.ca/ca/en/browse/90155046/agarose-electrophoresis-gels?page=1 [40] Invitrogen, "SYBR™ Safe DNA Gel Stain," Thermo Fisher Scientific, 2025. Available: https://www.thermofisher.com/order/catalog/product/S33102 [41] Thermo Scientific, "DNA Gel Loading Dye (6X)," Thermo Fisher Scientific, 2025. Available: https://www.thermofisher.com/order/catalog/product/R0611 [42] Invitrogen, "PureLink™ PCR Purification Kit," Thermo Fisher Scientific, 2025. Available: https://www.thermofisher.com/order/catalog/product/K310001 [43] Thermo Scientific, "T4 DNA Ligase (5 U/μL)," Thermo Fisher Scientific, 2025. Available: https://www.thermofisher.com/order/catalog/product/EL0011 [44] New England Biolabs, "T4 DNA Ligase Reaction Buffer," New England Biolabs, 2025. Available: https://www.neb.com/en-ca/products/b0202-t4-dna-ligase-reaction-buffer [45] Invitrogen, "Nuclease-Free Water (not DEPC-Treated)," Thermo Fisher Scientific, 2025. Available: https://www.thermofisher.com/order/catalog/product/AM9937 [46] Thermo Fisher Scientific, "LB Broth and LB Agar," Thermo Fisher Scientific, 2025. Available: https://www.thermofisher.com/ca/en/home/life-science/cell-culture/microbiological-culture/bacterial-growth-media/lb-broth-and-lb-agar.html